In order to train a computer vision related deep learning model I had to write a PyTorch custom dataloader for loading a set of annotation data. The data points were stored in JSON format and believe me, that massive JSON file was nearly 4GB! It was not a simple data structure with keys and values, but had a mixed set of data structures including lists, single float values and keys in String format.
How to use this format for your machine learning model? How can I train a model with data stored in the HDF5 format? We'll be studying the Hierarchical Data Format, as the data format is called, as well as how to access such files in Python - with h5py. Then, we actually create a Keras model that is trained with MNIST data, but this time not loaded from the Keras Datasets module - but from HDF5 files instead.
This is a format developed by the digitalsurf company to handle various types of scientific measurements data such as profilometer,SEM,AFM,RGB images, multilayer surfaces and profiles. Even though it is essentially a surfaces format, 1D signals are supported for spectra and spectral maps. Specifically, this file format is used by Attolight SA for the its Scanning Electron Microscope Cathodoluminescence (SEM-CL) hyperspectral maps. Metadata parsing is supported, including user-specific metadata, as well as the loading of files containing multiple objects packed together. In addition to image classification datasets, Caffe also have "HDF5Data" layer for arbitrary inputs. This layer requires all training/validation data to be stored in hdf5 format files.
HDF4 is the older version of the format, although still actively supported by The HDF Group. It supports a proliferation of different data models, including multidimensional arrays, raster images, and tables. Each defines a specific aggregate data type and provides an API for reading, writing, and organizing the data and metadata.
New data models can be added by the HDF developers or users. Nexus files can contain multiple datasets within the same file but the ordering of datasets can vary depending on the setup of an experiment or processing step when the data was collected. For example in one experiment Fe, Ca, P, Pb were collected but in the next experiment Ca, P, K, Fe, Pb were collected. HyperSpy supports reading in one or more datasets and returns a list of signals but in this example case the indexing is different.
To control which data or metadata is loaded and in what order some additional loading arguments are provided. HyperSpy can read and write the blockfile format from NanoMegas ASTAR software. It is used to store a series of diffraction patterns from scanning precession electron diffraction measurements, with a limited set of metadata. The header of the blockfile contains information about centering and distortions of the diffraction patterns, but is not applied to the signal during reading. Blockfiles only support data values of typenp.uint8 (integers in range 0-255).
Finally, one aspect that makes HDF5 particularly useful is that you can read and write files from just about every platform. The IDL language has supported HDF5 for years; MATLAB has similar support and now even uses HDF5 as the default format for its ".mat" save files. Bindings are also available for Python, C++, Java, .NET, and LabView, among others. Institutional users include NASA's Earth Observing System, whose "EOS5" format is an application format on top of the HDF5 container, as in the much simpler example earlier.
Even the newest version of the competing NetCDF format, NetCDF4, is implemented using HDF5 groups, datasets, and attributes. You have seen that using the .h5 format is simple and clean as it only creates one single file, whereas using tensorflow native format creates multiple folders and files, which is difficult to read. So, you might be thinking that why should we use tensorflow native format?
The answer to this is that in the TensorFlow native format, everything is structural and organized in its place. For example, the .pb file contains structural data that can be loaded by multiple languages. Some of the advantages of TF native format are listed in the following.
This results in a truly hierarchical, filesystem-like data format. In fact, resources in an HDF5 file can be accessed using the POSIX-like syntax /path/to/resource. Metadata is stored in the form of user-defined, named attributes attached to groups and datasets. More complex storage APIs representing images and tables can then be built up using datasets, groups and attributes. The computation graph of custom objects such as custom layers is not included in the saved file.
At loading time, Keras will need access to the Python classes/functions of these objects in order to reconstruct the model. Consider the multi-terabyte datasets that can be sliced as if they were real NumPy arrays. Thousands of datasets will be able to be stored in a single file and categorized. They can be tagged based on categories or however we want. H5Py can directly use NumPy and Python metaphors such as their NumPy array syntax and dictionary. For example datasets in a file can be iterated over and over or the attributes of the datasets such as .dtype or .shape can be checked out.
External losses & metrics added via model.add_loss()& model.add_metric() are not saved . If you have such losses & metrics on your model and you want to resume training, you need to add these losses back yourself after loading the model. Note that this does not apply to losses/metrics created inside layers viaself.add_loss() & self.add_metric().
As long as the layer gets loaded, these losses & metrics are kept, since they are part of the call method of the layer. Do note that there's also a different way of working with HDF5 files in Keras - being, with the HDF5Matrix util. While this works great, I found it difficult to adapt data when using it.
A global dataset cache is used to store the metadata information of a dataset. Given there are only a few pieces of information needed in order to know how to read a dataset, the H5Coro library maintains a large set of the most recently read datasets. Our initial architecture for processing ICESat-2 data in S3 used the native HDF5 library with the REST-VOL connector to read datasets via HDF5's Highly Scalable Data Service .
HSDS was run as a cluster of Docker containers on each EC2 instance we deployed to, and fanned out requests to S3 across multiple reader nodes. This provided a significant performance improvement over other approaches at the time, which included downloading the entire file or mounting the remote file for local access. Of the three methods, LMDB requires the most legwork when reading image files back out of memory, because of the serialization step. Let's walk through these functions that read a single image out for each of the three storage formats.
After training and saving the loading the model how I can predict the output of single data input ? Effectively I need to know how to use the predict function on the trained model. In scientific computing, sometimes, we need to store large amounts of data with quick access, the file formats we introduced before are not going to cut it.
You will soon find there are many cases, HDF5 is the solution. It is a powerful binary data format with no limit on the file size. It provides parallel IO (input/output), and carries out a bunch of low level optimizations under the hood to make the queries faster and storage requirements smaller. HyperSpy can read heater, biasing and gas cell log files for Protochips holder. The format stores all the captured data together with a small header in a csv file.
The reader extracts the measured quantity (e. g. temperature, pressure, current, voltage) along the time axis, as well as the notes saved during the experiment. The reader returns a list of signal with each signal corresponding to a quantity. Since there is a small fluctuation in the step of the time axis, the reader assumes that the step is constant and takes its mean, which is a good approximation. Further release of HyperSpy will read the time axis more precisely by supporting non-linear axis. The architecture of SlideRule cannot be built around a single dataset or file format, but must be easily expandable to handle datasets from other science communities stored in their native formats. First of all, all libraries support reading images from disk as .png files, as long as you convert them into NumPy arrays of the expected format.
This holds true for all the methods, and we have already seen above that it is relatively straightforward to read in images as arrays. Presumably, you have them already on disk somewhere, unlike our CIFAR example, so by using an alternate storage method, you are essentially making a copy of them, which also has to be stored. Doing so will give you huge performance benefits when you use the images, but you'll need to make sure you have enough disk space. In practice, the write time is often less critical than the read time. Imagine that you are training a deep neural network on images, and only half of your entire image dataset fits into RAM at once. Each epoch of training a network requires the entire dataset, and the model needs a few hundred epochs to converge.
You will essentially be reading half of the dataset into memory every epoch. Though this is a very simple data structure, you can expand this to complex and large files. You'll find it pretty easy to use HDF5 instead of using huge lists inside init of custom dataloaders. Here's a rough sketch of the PyTorch custom dataset class I created for the above example. In addition to these advances in the file format, HDF5 includes an improved type system, and dataspace objects which represent selections over dataset regions.
The API is also object-oriented with respect to datasets, groups, attributes, types, dataspaces and property lists. It lacks a clear object model, which makes continued support and improvement difficult. Supporting many different interface styles leads to a complex API.
Support for metadata depends on which interface is in use; SD objects support arbitrary named attributes, while other types only support predefined metadata. Perhaps most importantly, the use of 32-bit signed integers for addressing limits HDF4 files to a maximum of 2 GB, which is unacceptable in many modern scientific applications. The next question is, how can weights be saved and loaded to different models if the model architectures are quite different?
The solution is to use tf.train.Checkpoint to save and restore the exact layers/variables. SavedModel is the more comprehensive save format that saves the model architecture, weights, and the traced Tensorflow subgraphs of the call functions. This enables Keras to restore both built-in layers as well as custom objects.
In this blog post, we answered the question how to use datasets represented in HDF5 files for training your Keras model? H5 is a file format to store structured data, it's not a model by itself. Keras saves models in this format as it can easily store the weights and model configuration in a single file. It is an open-source file which comes in handy to store large amount of data. As the name suggests, it stores data in a hierarchical structure within a single file. So if we want to quickly access a particular part of the file rather than the whole file, we can easily do that using HDF5.
This functionality is not seen in normal text files hence HDF5 is becoming seemingly popular in fact of being a new concept. One important feature is that it can attach metaset to every data in the file thus provides powerful searching and accessing. Let's get started with installing HDF5 to the computer.
This is the file format used by the software package Element Identification for the Thermo Fisher Scientific Phenom desktop SEM. It is a proprietary binary format which can contain images, single EDS spectra, 1D line scan EDS spectra and 2D EDS spectrum maps. The reader will convert all signals and its metadata into hyperspy signals.
Signals saved as nxs by this plugin can be loaded normally and the original_metadata, signal data, axes, metadata and learning_results will be restored. To preserve the signal details an additional navigation attribute is added to each axis to indicate if is a navigation axis. One word of warning - don't use HDF5 if you are storing your data on S3. Because S3 treats objects as opaque blobs you will need to download the entire file to local storage every time you want to read it, even if you only want to read one tiny table from a 100GB file. If you are using S3 you are far better off saving each table in a separate parquet file and the metadata in JSON files. This way you can access the tables directly from S3 using Spark or Dask.
The H5Coro library was written to optimize access to the ICESat-2 H5 datasets. These datasets are typically numerical time-series which are chunked, compressed, and stored sequentially in memory. This layout is exploited in order to quickly determine the start and stop location of the data to be read, and works well when subsetting ICESat-2 data.
As a result, the implementation focuses on the most efficient way to retrieve the values of a dataset and nothing else. A metadata repository is needed to hold pointers into the original H5 files which HSDS uses to know how to read the various datasets in the file. This means that before any H5 file in S3 can be read by our system, it must first be loaded through an HSDS pipeline to build and store the metadata for it. A typical region of interest consisting of 60 granules can take over 8 hours to load using a single c5.xlarge EC2 instance.
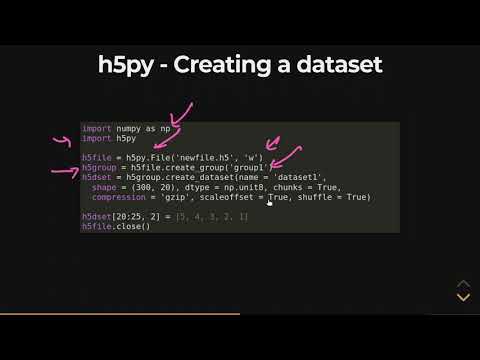
Our code opens the HDF5 we wrote above, prints the HDF5 attributes from the file, reads the two datasets, and then prints them out as columns. Of course, real code might add some error-handling and extracting other useful stuff from the file. In addition to the MEX format, we also provide matrices in the Hierarchical Data Format . H5 is a binary format that can compress and access data much more efficiently than text formats such as MEX, which is especially useful when dealing with large datasets. You've seen evidence of how various storage methods can drastically affect read and write time, as well as a few pros and cons of the three methods considered in this article. While storing images as .png files may be the most intuitive, there are large performance benefits to considering methods such as HDF5 or LMDB.






















No comments:
Post a Comment
Note: Only a member of this blog may post a comment.